Using conda environments¶
Let's get started with conda!
To follow along with this lesson, we have included several short video demonstrations of the steps described in this tutorial. We are using a binder with an Rstudio interface. Binders use collections of files from Github repositories with instructions on software installation to create small (and free!) computing environments. They are used for teaching and demonstrating software functionality or analysis workflows.
Open the binder for this lesson in a new tab (i.e., by typing Ctrl and clicking link): Click me to launch binder!
It may take 3-4 minutes for the binder to load!
Info
For this lesson, we are using Rstudio to teach you conda because it consolidates showing the conda commands, terminal, and file system all on 1 screen. In practice, you can use conda through a command-line terminal interface without Rstudio.
We are using 3 of the Rstudio panels for this lesson: Source panel to run conda commands, Terminal panel to execute code, and File panel to view input/output files. You can rearrange and minimize unused panels to help with viewing, as shown in the video below:
Warning
What happens if I get a 502, 503, or 504 error from the binder?
Try clicking on the launch button again to re-launch. The binder or internet connection may have timed out.
Conda is already installed in the binder so the next step is to set it up. We'll talk more about setting conda up on your local system later in the lesson!
Initialize conda¶
To follow along, copy/paste commands into the terminal OR run the commands from the "workshop_commands.sh" file in the binder (in File Rstudio panel). Either click Run or type Cmd+Enter on Macs and Ctrl+Enter on Windows computers.
The conda installer sets up two things: Conda and the base environment (also called "root"). The base environment contains a version of python (specified during installation) and some basic packages. As illustrated below, you can then create additional environments with their own software installations, including other versions of the same software (i.e., python 3 in base environment and python 2.7 in a separate environment).
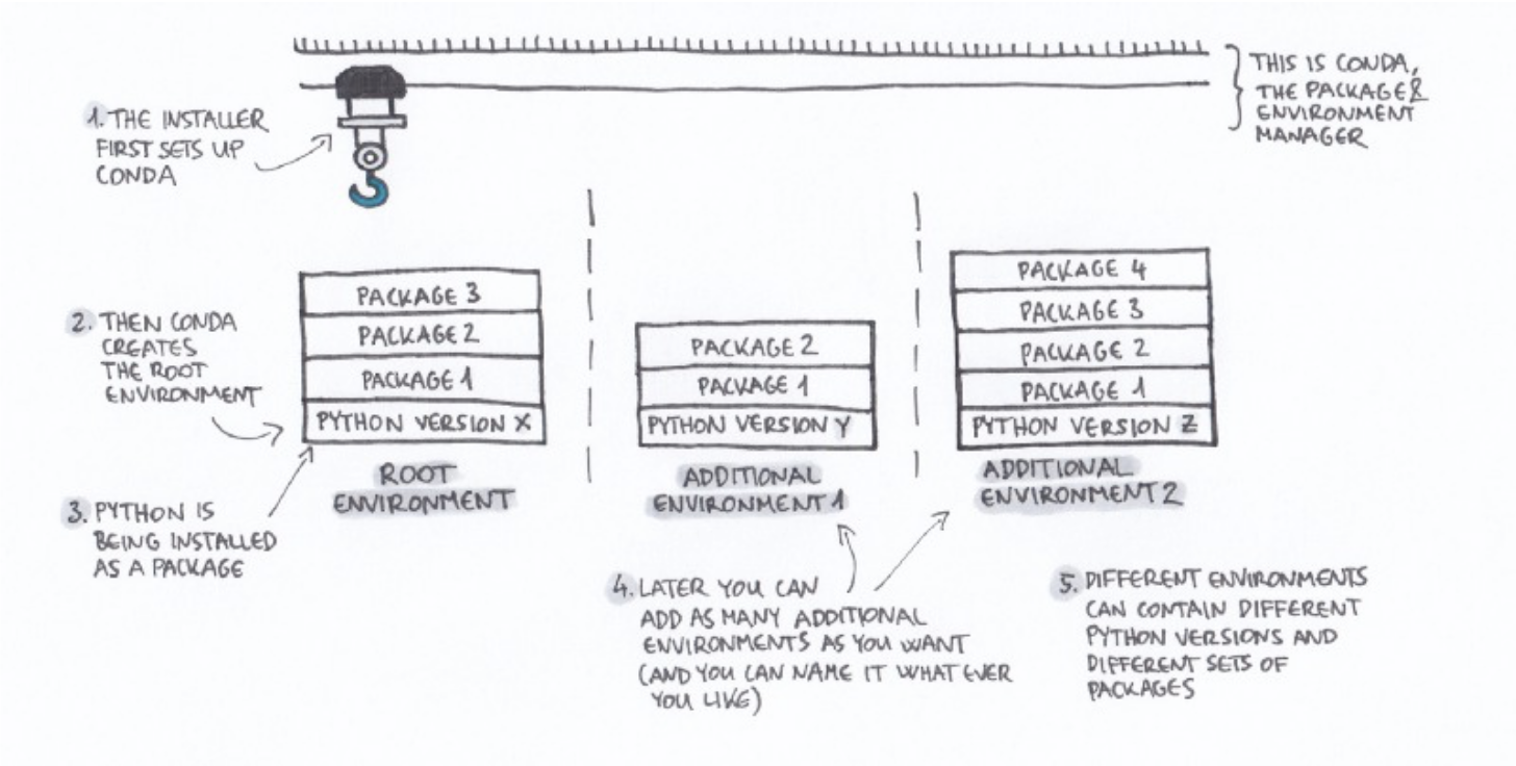
Image credit: Gergely Szerovay
Setup the conda installer and initialize the settings:
conda init
The binder auto-generates a very long command prompt. We will shorten it to $:
echo "PS1='\w $ '" >> .bashrc
Re-start terminal for the changes to take effect (type exit and then open a new terminal). These steps are demonstrated in the video below:
We are currently in the "base" conda environment (Note: the binder's terminal will not show (base) in the command prompt until we deactivate an environment. Your own conda installation should show (base) after initialization).
Conda channels: Searching for software¶
The channels are places where conda looks for software packages. The default channel after conda installation is set to Anaconda Inc's channels (Conda's Developer). Below, we demonstrate how to list the binder's default channels and the (base) environment packages, after which we add new channels and set channel priority:
conda config --show channels
# get list of packages in base environment
# shows channel, package version, and build number for each package
# information for specifying packages (specific versions, builds, or default (latest))
conda list | less -S
Note
You might notice that our installation of conda on the binder already had the defaults and conda-forge channels. This is due to the binder's set up. But in practice on your own system, it's important to add the channels as shown in this lesson.
Channels exist in a hierarchical order. By default, conda searches for packages based on:
Channel priority > package version > package build number
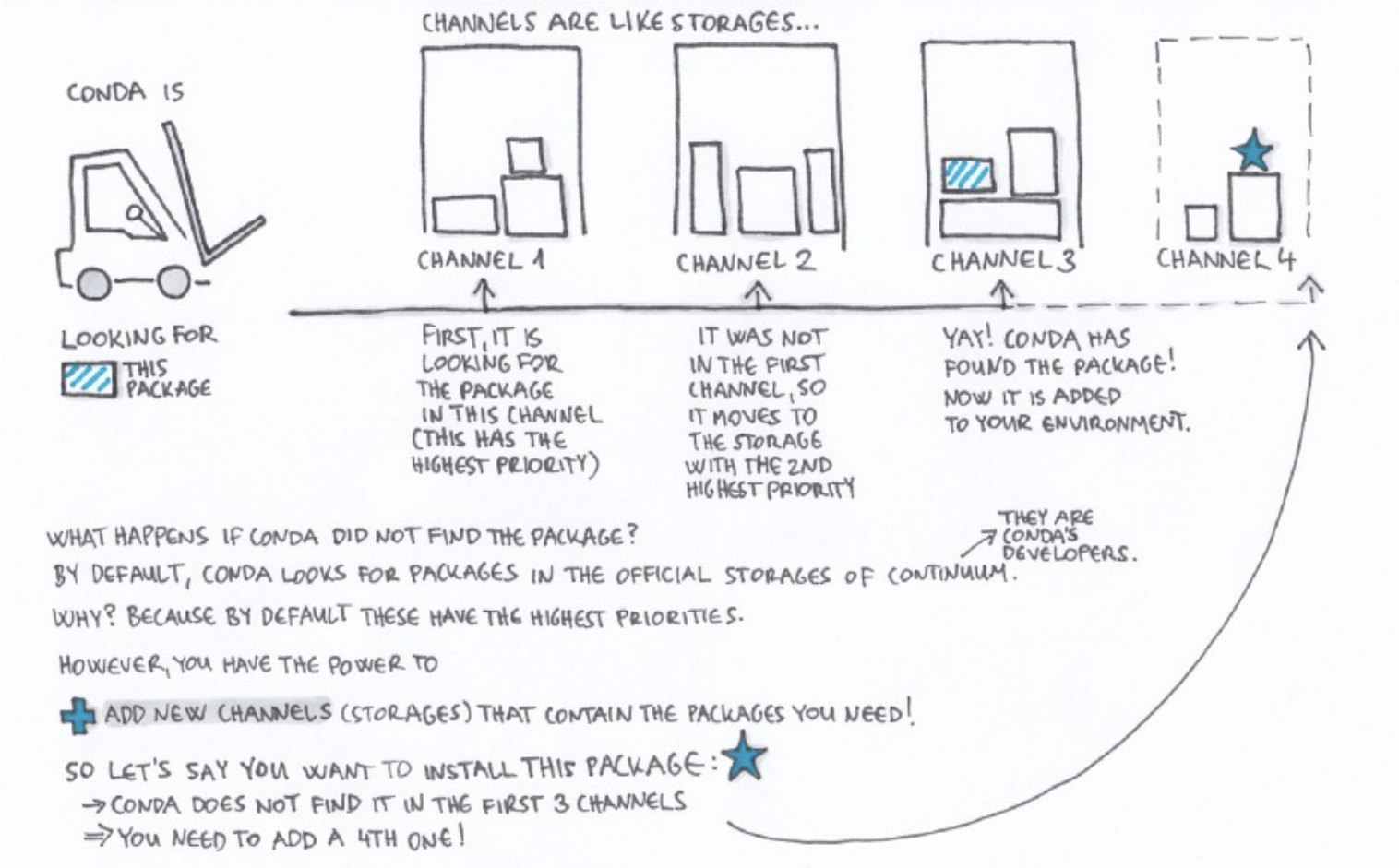
Image credit: Gergely Szerovay
Info
Commonly used channels:
- In the absence of other channels, conda searches the
defaultsrepository conda-forgeandbiocondaare channels that contain community-contributed softwareBiocondaspecializes in bioinformatics softwareBiocondasupports only 64-bit Linux and Mac OS- package list
conda-forgecontains many dependency packages- You can even install R packages with conda!
We will update the channel list order and add bioconda since we are using bioinformatics tools today. The order of the channels matters!
First, add the defaults channel with the conda config --add channels command. We can check the channel priority order with the conda config --get channels command.
conda config --add channels defaults
conda config --get channels
Then add the bioconda channel:
conda config --add channels bioconda
conda config --get channels
Lastly, add the conda-forge channel to move it to top of the list, following Bioconda's recommended channel order. This is because many packages on bioconda rely on dependencies that are available on conda-forge, so we want conda to search for those dependencies before trying to install any bioinformatics software.
conda config --add channels conda-forge
conda config --get channels
With this configuration, conda will search for packages first in conda-forge, then bioconda, and then defaults.
Info
Another way to add channels is:
conda config --prepend channels bioconda
Install Software and Create Environments¶
For our demo, we need to install FastQC, a commonly used software tool that provides quality control reports for raw sequencing data.
Search for software (fastqc):
conda search fastqc
It may take a few seconds for the software list to display. The table shows all the versions and builds of fastqc available for installation with conda. They are all stored in the bioconda channel.
Loading channels: done
# Name Version Build Channel
fastqc 0.10.1 0 bioconda
fastqc 0.10.1 1 bioconda
fastqc 0.11.2 1 bioconda
fastqc 0.11.2 pl5.22.0_0 bioconda
fastqc 0.11.3 0 bioconda
fastqc 0.11.3 1 bioconda
fastqc 0.11.4 0 bioconda
fastqc 0.11.4 1 bioconda
fastqc 0.11.4 2 bioconda
fastqc 0.11.5 1 bioconda
fastqc 0.11.5 4 bioconda
fastqc 0.11.5 pl5.22.0_2 bioconda
fastqc 0.11.5 pl5.22.0_3 bioconda
fastqc 0.11.6 2 bioconda
fastqc 0.11.6 pl5.22.0_0 bioconda
fastqc 0.11.6 pl5.22.0_1 bioconda
fastqc 0.11.7 4 bioconda
fastqc 0.11.7 5 bioconda
fastqc 0.11.7 6 bioconda
fastqc 0.11.7 pl5.22.0_0 bioconda
fastqc 0.11.7 pl5.22.0_2 bioconda
fastqc 0.11.8 0 bioconda
fastqc 0.11.8 1 bioconda
fastqc 0.11.8 2 bioconda
fastqc 0.11.9 0 bioconda
Now, let's create a conda environment with fastqc installed in it, as demonstrated below:
Create conda environment and install FastQC. This takes a few minutes (you'll see the message "Solving environment").
The -y flag tells conda not to ask you for confirmation about downloading software. The --name (or -n) flag specifies the environment's name. The last element of the command, fastqc, specifies the software package to install.
conda create -y --name fqc fastqc
More options to customize the environment are documented under the help page for this command: conda create -h.
The software you installed will only be available to use after you activate the environment:
conda activate fqc
This command shows you information about the activated conda environment:
conda info
One way to make sure the software works is to check the version:
fastqc --version
Info
To go back to (base) ~ $ environment:
conda deactivate
High-throughput sequencing data quality control steps often involve FastQC and Trimmomatic. Trimmomatic is useful for read trimming (i.e., adapters). There are multiple ways we could create a conda environment that contains both software programs:
Method 1: install software in existing environment¶
We could add trimmomatic to the fqc environment:
conda install -y trimmomatic=0.36
conda list # check installed software
We can specify the exact software version with = and a version number. The default is to install the latest version, but sometimes your workflow may depend on an older version.
Info
Software can also be installed by specifying the channel with -c flag:
conda install -c conda-forge -c bioconda trimmomatic=0.36
or if needed, by specifying version and build (the default is to install the latest version and build):
conda install trimmomatic=0.32=0
When you switch conda environments, conda changes the file path (and other environment variables) to searches for software packages in different folders.
Let's check the PATH for method 1:
echo $PATH
You should see that the first element (/srv/conda/envs/fqc/bin:) in the file path changes each time you switch environments!
/srv/conda/envs/fqc/bin:/srv/conda/condabin:/srv/conda/envs/notebook/bin:/srv/conda/condabin:/home/jovyan/.local/bin:/home/jovyan/.local/bin:/srv/conda/envs/notebook/bin:/srv/conda/bin:/srv/npm/bin:/usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin
Method 2: install both software during environment creation¶
For this method, we list trimmomatic=0.36 after fastqc to create an environment with both installed, all with 1 command. Like above, remember to activate the environment and then you can check the list of packages to verify installation and check the PATH to verify that conda switched to the fqc_trim directory.
conda deactivate
conda create -y --name fqc_trim fastqc trimmomatic=0.36
conda activate fqc_trim
# check installed software
conda list
# path for method 2
echo $PATH
The following methods use an external file to specify the packages to install:
Method 3: specify software to install with a YAML file¶
Often, it's easier to create environments and install software using a YAML file that specifies all the software to be installed as demonstrated in the video. For our example, we are using a file called test.yml.
Let's start back in the (base) environment.
conda deactivate
The test.yml file contains the following in YAML format:
name: qc_yaml #this specifies environment name
channels:
- conda-forge
- bioconda
- defaults
dependencies:
- fastqc
- trimmomatic=0.36
Info
YAML is a file format that is easy for both computers and humans to read. The YAML file extension is .yml and these files can be generated in any text editor.
For conda, the name: is optional (it can also be specified in the conda env create command), but it must have a list of channels: and a list of dependencies:. Notice that the channels are list with highest to lowest priority.
Create the environment - note the difference in conda syntax. This method uses the conda env create command instead of conda create. The -f (or --file) flag specifies the file with the channels and software to set up.
# since environment name specified in yml file, we do not need to use -n flag here
conda env create -f test.yml
conda activate qc_yaml
# check installed software
conda list
Info
Note that you can export conda environments as YAML files as well. The --from-history option outputs only the packages you've installed in this environment, as opposed to all the other default packages in the environment.
conda activate qc_yaml
conda env export --from-history > qc_env.yml
conda deactivate
Method 4: Install exact environment¶
For this approach, we export a list of the exact software package versions installed in a given environment and use it to set up new environments. This set up method won't necessarily install the latest version of a given program, but it will replicate the exact environment set up you exported from.
conda activate fqc
conda list --export > packages.txt
conda deactivate
Two options -
1) install the exact package list into an existing environment:
conda install --file=packages.txt
OR
2) set up a new environment with the exact package list:
conda env create --name qc_file -f packages.txt
Managing Environments¶
At this point, we have several conda environments! To see a list:
conda env list
The current environment you're in is marked with an asterisk *.
Note
There are a few redundant commands in conda. For example, this command does exactly the same thing as the one above:
conda info --envs
Generally, you want to avoid installing too many software packages in one environment. The more software you install, the longer it takes for conda to resolve compatible software versions for an environment (it'll take longer and longer at the "Solving environment" stage).
For this reason, and in practice, people often manage software for their workflows with multiple conda environments.
Running FastQC in a conda environment¶
Let's run a small analysis with FastQC in the fqc environment we created above:
If not already done, activate one of the environments we created, e.g.,:
conda activate fqc
Let's make sure the software was installed correctly by looking at the help documentation:
fastqc --help
Output should look like:
FastQC - A high throughput sequence QC analysis tool
SYNOPSIS
fastqc seqfile1 seqfile2 .. seqfileN
fastqc [-o output dir] [--(no)extract] [-f fastq|bam|sam]
[-c contaminant file] seqfile1 .. seqfileN
...
Download data (a yeast sequence file):
curl -L https://osf.io/5daup/download -o ERR458493.fastq.gz
Check out the data:
The gunzip -c command allows us to see the unzipped version of the file without actually unzipping it (you can verify this by checking the file extension after running this command!). The | is called a pipe and it takes the output of the gunzip -c command and hands it to the wc word count command. The -l flag tells wc we want to count the number of lines in the file.
gunzip -c ERR458493.fastq.gz | wc -l
There should be 4,375,828 lines in the file.
What does the fastq file look like?
Here are two ways to look at the sequence read file:
1. Use the gunzip -c and pipe the output to the head command to show the first 10 lines of the file:
gunzip -c ERR458493.fastq.gz | head
2. Use the less command to scroll through the file:
less ERR458493.fastq.gz
The beginning of the fastq format sequence file should look like this, where the 1st line is the sequence read ID (starts with @), the 2nd line is the DNA sequence, the 3rd is sequence separator +, and the 4th is the Phred quality score associated with each base pair in ASCII format.
@ERR458493.1 DHKW5DQ1:219:D0PT7ACXX:1:1101:1724:2080/1
CGCAAGACAAGGCCCAAACGAGAGATTGAGCCCAATCGGCAGTGTAGTGAA
+
B@@FFFFFHHHGHJJJJJJIJJGIGIIIGI9DGGIIIEIGIIFHHGGHJIB
@ERR458493.2 DHKW5DQ1:219:D0PT7ACXX:1:1101:2179:2231/1
ACTAATCATCAACAAAACAATGCAATTCAAGACCATCGTCGCTGCCTTCGC
+
B@=DDFFFHHHHHJJJJIJJJJJJIJJJJJJJJJJJJJJJJJJJJIJJJJI
@ERR458493.3 DHKW5DQ1:219:D0PT7ACXX:1:1101:2428:2116/1
CTCAAAACGCCTACTTGAAGGCTTCTGGTGCTTTCACCGGTGAAAACTCCG
...
If you used the less command, type Q to exit the page.
Run FastQC!
fastqc ERR458493.fastq.gz
On the terminal screen, FastQC prints analysis progress:
Started analysis of ERR458493.fastq.gz
Approx 5% complete for ERR458493.fastq.gz
Approx 10% complete for ERR458493.fastq.gz
Approx 15% complete for ERR458493.fastq.gz
Approx 20% complete for ERR458493.fastq.gz
Approx 25% complete for ERR458493.fastq.gz
Approx 30% complete for ERR458493.fastq.gz
Approx 35% complete for ERR458493.fastq.gz
...
Analysis complete for ERR458493.fastq.gz
The final output file is called "ERR458493_fastqc.html".
You can click on the .html file in the File panel to open it in a web browser. This is the quality check report for our yeast sequence file.