Run the whole pipeline!¶
Step 11: Running the pipeline¶
Once you've decorated the entire Snakefile, you should be able to run through the whole workflow using either the rule name:
snakemake -p make_vcf -j 2
or by using the file name:
snakemake -p variants.vcf -j 2
Recollect that Snakemake will execute the first rule in the Snakefile by default. We can use this feature by creating a default rule called all at the top of the Snakefile:
rule all:
input: "variants.vcf"
and run:
snakemake -p all -j 2
This rule runs through the entire workflow with a single command! This is much better than running each command one by one!
The final snakemake workflow looks like this:
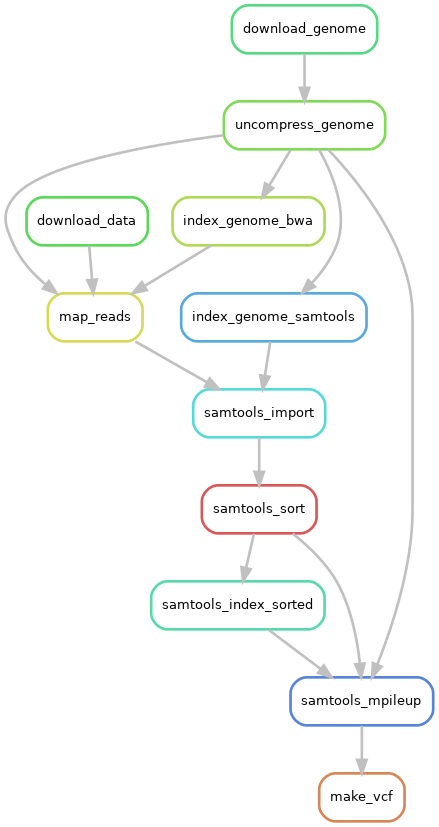
Note
The diagram above is called a directed acyclic graph (DAG) and it is how snakemake interprets the workflow from the Snakefile. You can read more about the syntax information on the snakemake documentation.
To generate a DAG file, we need Graphviz, a graph visualization software, and dot to create directed graphs. We can use conda to create a new environment with the required packages.
# initialize and reset the bash with conda
conda init bash
source .bashrc
# create a new environment and install Graphviz
conda create -y -n graphs graphviz
# activate conda environment
conda activate graphs
# create dag png file
snakemake --dag variants.vcf | dot -Tpng > vc_workflow_dag.png
Step 12: Looking at VCF files¶
Finally, let's look at the output!
less variants.vcf
We can look at the alignment by running the alignment viewer in samtools:
samtools tview -p ecoli:4202391 SRR2584857.sorted.bam ecoli-rel606.fa
Navigating samtools tview
- Access the help menu in
samtools tviewby hittingshiftkey and?key. - Exit help menu by hitting
qkey - Go to a specific position location in the alignment by hitting
/key.
Conclusion¶
Exit (snaketest) conda environment to return to (base) environment:
conda deactivate
Hopefully, you have now:
- learned how to write basic workflows with Snakemake rules
- learned variable substitution for Snakemake rules
- learned wildcard matching for Snakemake rules
- understood why workflow systems can help you do your computing more easily!
Key Points
- Snakefile defines a Snakemake workflow
- the rules specify steps in the workflow
- at the moment (and in general), they run shell commands
- you can "decorate" the rules to link the dependencies between rules
- basic decoration comes in the form of
input:andoutput:which are lists of one or more files, quoted, separated by commas - the rules are connected by matching filenames
- tabs are important syntax feature in Snakemake